
Andrea Pauli
former member
About
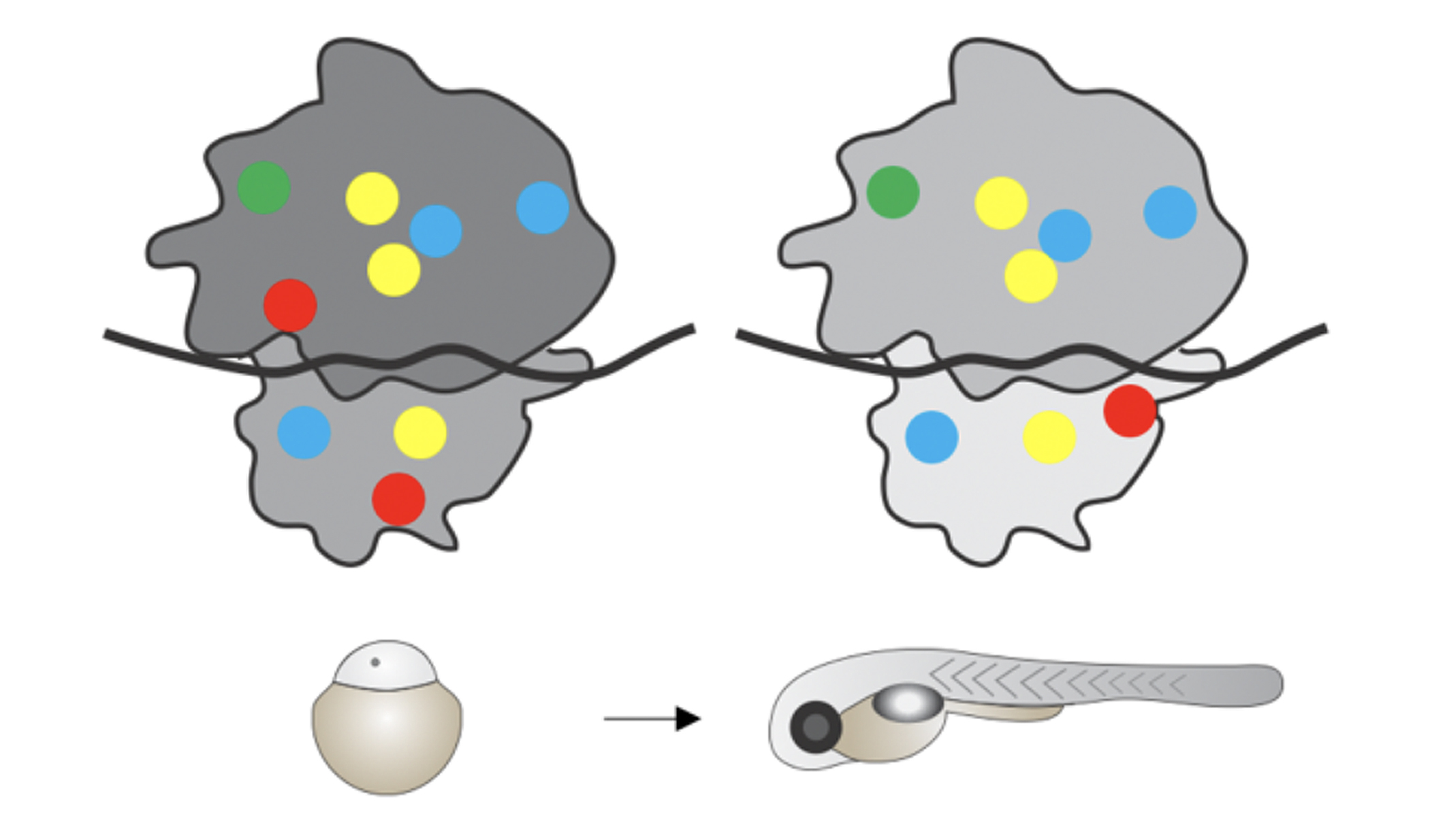
Dynamics of rRNA modifications during the oocyte-to-embryo transition
Life of sexually reproducing organisms starts with the fusion of two highly specialized cells, the egg and the sperm, to form a single cell, the zygote. This totipotent cell gives rise to all cells of the future organism. Triggered by two key discoveries, namely that i) translation is more wide-spread than anticipated and often occurs outside of known, annotated genes, and ii) that some of these newly identified translated regions can encode previously missed, essential small proteins (Pauli & Valen et al., 2012; Chew et al., 2013; Pauli et al., 2014; Chew et al., 2016; Cabrera-Quio & Herberg, Pauli, 2016), our main focus over the past years has been on the functional characterization of novel, essential short proteins during embryogenesis (Pauli et al., 2014; Herberg et al., 2018). In addition, stemming from our general interest in regulatory principles that govern the large-scale transformation of the egg cytoplasm into the cytoplasm of metabolically active, embryonic cells, we are investigating the underlying principles of translational regulation during the oocyte-to-embryo transition. The long-term vision of the Pauli lab is to unravel new concepts and molecular principles governing the oocyte-to-embryo transition.
To address these questions, we are mainly using zebrafish embryos since they are a powerful model of vertebrate embryogenesis. We combine various approaches, ranging from genetics, molecular and cell biology, biochemistry and structural biology to live cell imaging, genomics and evolutionary analyses.
Within the SFB the Pauli group aims to focus on the dynamics and functional implications of rRNA modifications during embryogenesis.
Phone
+43 1 79730 3500
E-mail
andrea.pauli@imp.ac.at
Research Institute of Molecular Pathology (IMP)
Pauli Lab (3rd floor)
Campus-Vienna-Biocenter 1 A-1030 Vienna, Austria
